Abstract
The complete chloroplast genome (cpDNA) sequence of Aconitum volubile var. pubescens, an im- portant and popular traditional medicinal plant species for Aconiti Kusnezoffii Tuber, is determined in this study. The cpDNA of A. volubile var. pubescens was 155,872 bp in length with 38.12% GC content that composed an 86,348 bp large single-copy (LSC), a 16,944 bp small single-copy (SSC), and a pair of 26,290 bp inverted repeats (IRA and IRB). The cpDNA of A. volubile var. pubescens encodes 131 predicted unique genes including 86 protein-encoding (CDS) genes, 8 ribosomal RNA (rRNA) genes and 37 transfer RNA (tRNA) genes. Phylogenetic analysis revealed that the complete cpDNA sequence could be applicable as a super barcode to discriminate A. volubile var. pubescens from its closely related species and provides diverse information to investigate molecular DNA markers for the Aconiti Kusnezoffii Tuber.
Introduction
Aconitum is a large eudicot genus consisted approximately 300 species and an important medic- inal plant resources for removing pain, anti-inflammation, paralyzing a part of body etc. in oriental traditional medicines1). In national pharmacopoeia of Korea and China, it is differentially described that the original plant species of Aconiti Kusnezoffii Tuber, namely Cho-o in Korean and Cao-wu in Chinese, is defined as the tuber of Aconitum kusnezoffii Reichb., Aconitum ciliare DC. (a syno- nym of A. volubile var. pubescens Regel), and Aconitum jaluense var. triphyllum (Nakai) U.C.La (a synonym of Aconitum triphyllum Nakai) in Korea Herbal Pharmacopoeia but it is defined as the tuber of only A. kusnezoffii in Chinese Pharmacopoeia2). Furthermore, the root of Aconitum carmichaeli Debaeux is described as additional herbal medicine, namely Aconiti Lateralis Radix
The complete chloroplast genome sequence of a medicinal plant Aconitum volubile var. pubescens Regel
(Ranunculaceae)
Yang Sungyu, Kim Wook-jin, Park Inkyu, Yoe Sang-min, Moon Byeong-cheol* K-herb Research Center, Korea Institute of Oriental Medicine, Daejeon, Republic of Korea
*Correspondence: Moon Byeong-cheol Tel: +82-42-868-9530 Fax: +82-42-863-9434 E-mail: bcmoon@kiom.re.kr)
· Received 2016-10-17, accepted 2016-10-25, online-published 2016-10-25.
Keywords: Chloroplast genome, Aconitum, Aconitum volubile var. pubescens, medicinal plant, Aconiti Kusnezoffii Tuber, Ranunculaceae
(Bu-ja in Korean and Fu-zi in Chinese), in both country for some different clinical purpose with Aconiti Kusnezoffii Tuber2). Since the morphological similarity of the tuberous root and aerial part among the several species, however, the precise identification of the species is very difficult.
Therefore, the information useful for developing DNA based molecular marker such as sequences of diverse DNA barcode regions and super barcodes including complete plastid genome sequences because these herbal materials distributed as the dried root slices and its processed herbal drugs3-4).
Main Subject
It has been reported that DNA barcode sequences obtained several cpDNA fragments provide limited information for identifying the species in plantae5). In recent studies, it is reported that the complete chloroplast genome (cpDNA) sequences provide abundant information for molecular breeding, DNA barcoding, population genetics and molecular phylogenetics in plants6). Especially, cpDNA permit the limit of resolution for the identification of plant taxa in species levels by pro- viding super DNA barcode information against those of traditional barcodes such as rbcL and matK genes7). This being so, many of the complete cpDNA sequences have been analyzed for identifying the species of medicinal plants and improving the quality control technique of herbal medicines8). For the analysis of phylogenetic relationship and evaluation of identification ability of super bar- code between Aconitum and closely related plant species, we sequenced and compared complete cpDNA sequences.
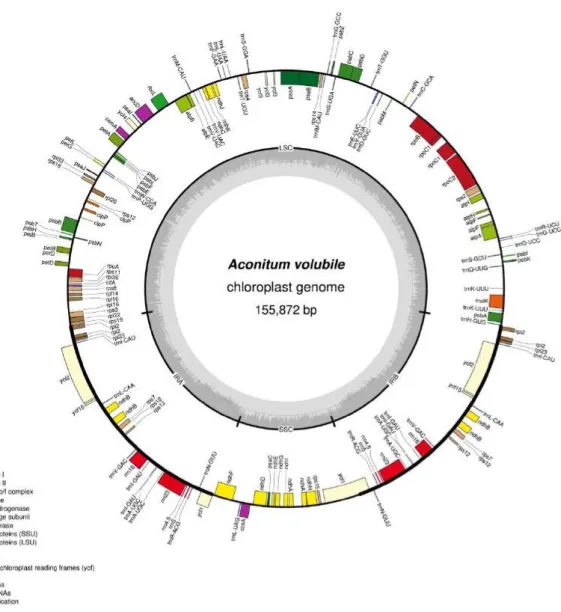
In this study, we sequenced cpDNA of A. volubile var. pubescens with Illumina whole genome next generation sequencing method. Depending on approximately 4.85 Gb raw sequences, we as- sembled the complete cpDNA sequence with de novo assembly of low coverage whole genome shotgun sequencing (dnaLCW) and annotated with DOGMA (Dual Organellar Genome Annotator)9). The cpDNA of A. volubile var. pubescens was 155,872 bp circular DNA (GenBank accession number KU556690) and similar in range with those from other angiosperms (Figure 1). The GC content was 38.12%, which was typical AT-reach and similar to those previously reported plant species10). It also showed a typical quadripartite structure and consisted a pair of IR regions, 26,290 bp IRA and IRB, separated by 86,348 bp LSC and 16,944 bp SSC. The complete cpDNA of A. volubile var. pubescens containing 131 functional genes consisted 80 protein-coding genes (CDS), 37 transfer RNA (tRNA) genes and 8 ribosomal RNA (rRNA) genes. Out of these genes, 8 protein-coding genes, 7 tRNA genes and 4 rRNA genes are duplicated in the IR regions and 14 and 3 genes contained a single intron and two introns, respectively.
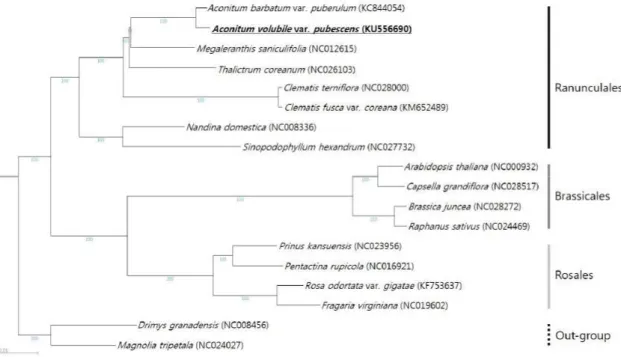
The phylogenetic tree was constructed to infer the molecular phylogenetic relationships and to
Figure 1. Gene map of the Aconitum volubile var. pubescens chloroplast genome. Genes drawn inside of the circle are transcribed clockwise, and those outside are counterclockwise. Genes belonging to different func- tional groups are color-coded. The darker gray in the inner circle corresponds to GC content, while the lighter gray corresponds to AT content.
Figure 2. Phylogenetic relationship of Aconitum volubile var. pubescens with other plant species belonging to the eudicots base on complete chloroplast genome sequences. Numbers in the nodes are the bootstrap values from 1000 replicates. The chloroplast genome sequences of Drimys granadensis and Magnolia tripe- tala was set as outgroup.
Conclusion
In this study, we sequenced the complete cpDNA for A. volubile var. pubescens, an important herbaceous plant species in Korea, using the Illumina Miseq sequencing method. Our results sug- gest that the cpDNA can be used as a super DNA barcode to distinguish A. volubile var. pubescens from its closely related species.
Acknowledgements
We thank the ‘Classification and Identification committee of the KIOM’ for the identification of plant materials and the Herbarium of Korea Standard Herbal Resources (herbarium code KIOM) for the provision of plant materials. This work was supported by a grant to the KIOM:
“Development of Foundational Techniques for the Domestic Production of Authentic Herbal
참고문헌
1. Dan B, Steven C, Erich S and Andrew G. Chinese Herbal Medicine Matria Medica. 3rd edition.
Washington:Eastland Press. 2004:673-81.
2. Korea Institute of Oriental Medicine. Defining Dictionary for Medicinal Herbs [Korean], Retrie ved October 10, 2016, from http://boncho.kiom.re.kr/codex/.
3. World Health Organization. Quality Control Methods for Medicinal Plant Materials, World Healt h Organization, Geneva. 1998.
4. Klich MA. Identification of common Aspergillus species. Centraalbureau Voor Schimmelculture s. Utrecht, The Netherlands. 2002.
5. Nock CJ, Waters DL, Edwards MA, Bowen SG, Rice N, Cordeiro GM and Henry RJ. Chloroplast genome sequences from total DNA for plant identification. Plant Biotechnology Journal. 2011;9:
328-33.
6. Nikiforova SV, Cavalieri D, Velasco R and Goremykin V. Phylogenetic analysis of 47 chloroplast genomes clarifies the contribution of wild species to the domesticated apple maternal line. Mol ecular Biological Evolution. 2013;30:1751 60.–
7. Friesen N, Pollner S, Bachmann K and Blattner FR. RAPDs and noncoding chloroplast DNA reveal a single origin of the cultivated Allium fistulosum from A. altaicum (Alliaceae). America n Journal of Botany. 1999;86:554 62.–
8. Liu Y, Huo N, Dong L, Wang Y, Zhang S, Young HA, Feng X and Gu YQ. Complete chloroplast genome sequences of Mongolia medicine Artemisia frigida and phylogenetic relationships with other plants. PLoS ONE. 2013;8(2):e57533. doi:10.1371/journal.pone.0057533.
9. Wyman SK, Jansen RK and Boor JL. Automatic annotation of organellar genomes with DOGMA.
Bioinformatics. 2004;20:3252-5.
10. Lu, D., Zhao, Y., Han, R, Wang, L and Qin P. The complete chloroplast genome sequence of the Purple Feathergrass Stipa purpurea (Poales: Poaceae), Conservation Genetic Resource s. 2016;8:101-4.