2019 년 1 학기
Text : Harper’s Biochemistry Refs: Devlin’s, Lippincott’s
Ch. 32. Nucleotides
2019 년 1 학기
Text : Harper’s Biochemistry Refs: Devlin’s, Lippincott’s
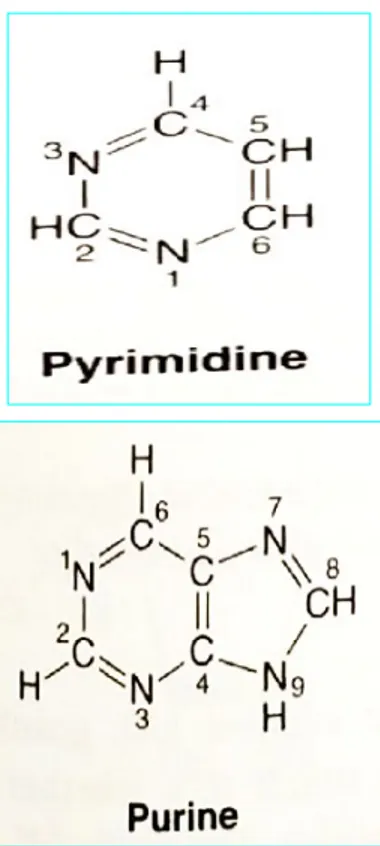
Figure 32-1 Purine and pyrimidine.
The atoms are numbered according to the international system.
Biomedical importance:
Monomeric unit of nucleic acids (precursors of nucleic acids) Metabolic functions
(energy metabolism, protein synthesis, regulation of enzyme activity, signal transduction) Coenzymes (vitamins and vitamin derivatives)
Donors & Acceptors of phosphate
Key biosynthetic intermediates of sugars & lipids Second messengers (cAMP & cGMP)
Chemotherapy of cancer, AIDS, & immune suppressors
•Nucleic acids:
P
hosphate(s) - (d)D-Ribose (sugar) - Base(s)(Purine/Pyrimidine)Figure 32-3 Ribonucleosides
Chemistry of purines, pyrimidines, nucleosides, & nucleotides
Base(s), Nucleosides, Nucleotides1) Base(s); ~ carbon/ nitrogen-containing heterocylic compounds
~ opposite directions of numbering six-atom rings (Fig. 32-1) 2) Nucleosides are N-Glycosides; (Fig. 32-3)
~ (d)D-Ribose (sugar) – Base(s)(Purine/Pyrimidine) ~ Sugar:
Ribonucleosides (D-ribose)
deoxyribonucleosides (2-deoxy-D-ribose)
~ Ribose(sugar) – Base(s)(Purine/Pyrimidine): -N-glycosidic bond,
Sugar - (N-1)pyrimidine, Sugar – (N-9)purine
•Nucleic acids: Phosphate(s) –
(d)D-Ribose (sugar) - Base(s)(Purine/Pyrimidine)
Deoxyadeno-sine
Figure 32-2 Tautomerism of the oxo and amino
functional groups of purines and pyrimidines
Physiological conditions Favor amino & oxo
Chemistry of purines, pyrimidines, nucleosides, & nucleotides
Base(s), Nucleosides, Nucleotides1) Base(s);
2) Nucleosides are N-Glycosides;
~ (d)D-Ribose (sugar) – Base(s)(Purine/Pyrimidine) 3) Nucleotides are phosphorylated nucleosides (Fig. 32-4)
~ In general, nucleosides with a phosphoryl group on the 5’-hydroxyl group of sugar
~ Additional phosphoryl group is ligated by acid anhydride bonds (high energy group transfer potential)
to the phosphoryl group of a mononucleotide, forming nucleoside diphosphate and triphos-phate (Fig. 32-4). --- AMP,, ADP,, ATP
•Nucleic acids:
Phosphate(s) – (d)D-Ribose (sugar) - Base(s)(Purine/Pyrimidine)
Chemistry of purines, pyrimidines, nucleosides, & nucleotides
Base(s), Nucleosides, Nucleotides1) Base(s);
2) Nucleosides are N-Glycosides;
~ (d)D-Ribose (sugar) – Base(s)(Purine/Pyrimidine) 3) Nucleotides are phosphorylated nucleosides (Fig. 32-4)
4) Heterocyclic N-Glycosides exist as Syn and Anti conformers (Fig. 32-5) * Anti conformers predominate.
Figure 32-5 The syn and
anti
conformers of adenosine dif -fer
with respect to orientation about the N-glycosidic bond
Steric hindrance by the heterocycle:
No freedom of rotation about the -N-glycosidic bond of nucleosides or nu-cleotides
Chemistry of purines, pyrimidines, nucleosides, & nucleotides
Base(s), Ribonucleosides, Ribonucleotides (Table 32-1. Fig. 32-6)•Nucleic acids:
Phosphate(s) – (d)D-Ribose (sugar) - Base(s)(Purine/Pyrimidine)
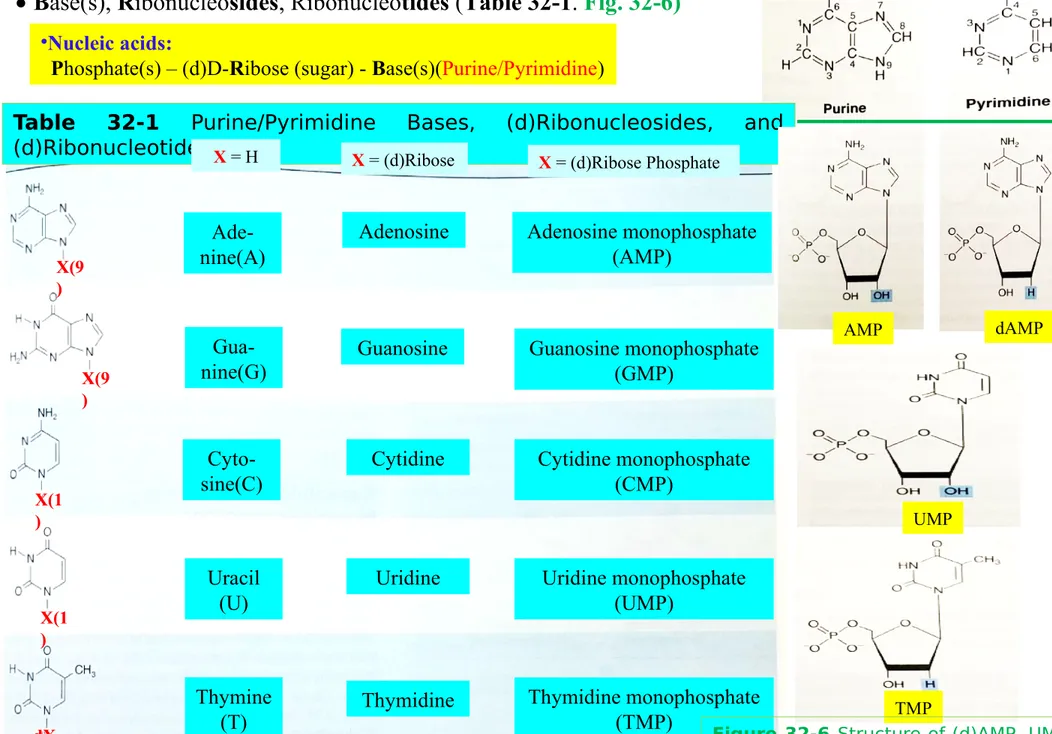
Table 32-1 Purine/Pyrimidine Bases, (d)Ribonucleosides, and
(d)Ribonucleotides Ade-nine(A) Gua-nine(G) Cyto-sine(C) X = H X(9 ) X(9 ) X(1 ) X(1 ) dX
X = (d)Ribose X = (d)Ribose Phosphate
Uracil (U) Thymine (T) Adenosine Cytidine Uridine Thymidine Guanosine
Figure 32-1 Purine and
pyrim-idine Adenosine monophosphate (AMP) Thymidine monophosphate (TMP) Guanosine monophosphate (GMP) Cytidine monophosphate (CMP) Uridine monophosphate (UMP) UMP TMP AMP dAMP
Figure 32-6 Structure of (d)AMP, UMP,
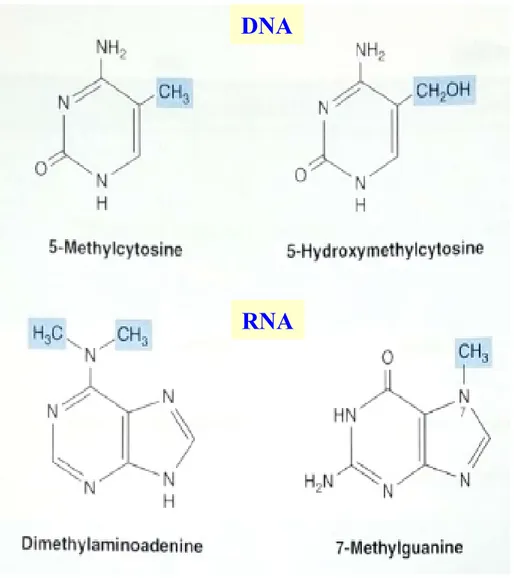
Figure 32-7 Four uncommon
but naturally occurring pyrimidines and purines Modification of polynucleotides can generate additional structures
Four uncommon (additional) bases in DNA and RNA (Fig. 32-7)
Figure 32-1 Purine and
pyrimi-dine DNA
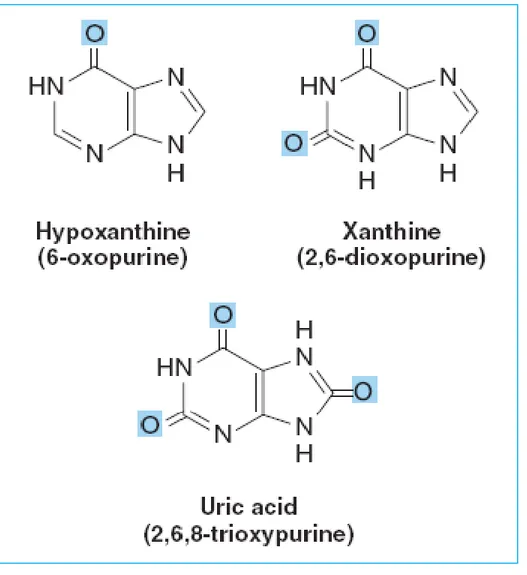
Figure 32-8 Structures of hypoxanthine, xanthine, and
uric acid
Modification of polynucleotides can generate additional structures Intermediates in the catabolism of adenine and guanine;
~ Hypoxanthine, Xanthine, Uric acid (Fig. 32-8) (Chapter 33)
Figure 32-1 Purine and
Fig. 32-9 Caffeine, a trimethylxanthine (coffee).
The dimethylxanthines theobromine (cocoa) and theo-phylline (tea) are similar but lack the methyl group at N-1 and at N-7, respectively.
Modification of polynucleotides can generate additional structures Methylated heterocycles (bases) of plants;
~ Caffeine (Xanthine derivatives; Fig. 32-8) (Fig. 32-9) Figure 32-1 Purine and pyrimi-dine
Caffeine
Nucleotides absorb ultraviolet light
~ Nucleotides absorb light at a wavelength at 260 nm
Fig. 32-10 cAMP, 3’,5’-cyclic AMP,
and cGMP, 3’,5’-cyclic GMP
Nucleotides serve diverse physiological functions
ATP (1 mmol/L): principal biological transducer of free energy Second messenger: cAMP (1 nmol/L) & (GTP) cGMP (Fig. 32-10)
Sulfate donor:
Adenosine-3'-phosphate-5-phosphosulfate (Fig. 32-11),
Figure 32-1 Purine and
pyrim-idine
Figure 32-11 Adenosine
3’-phos-phate-5’-phosphosulfate.
Methyl group donor:
S-Adenosylmethionine (Fig. 32-12)
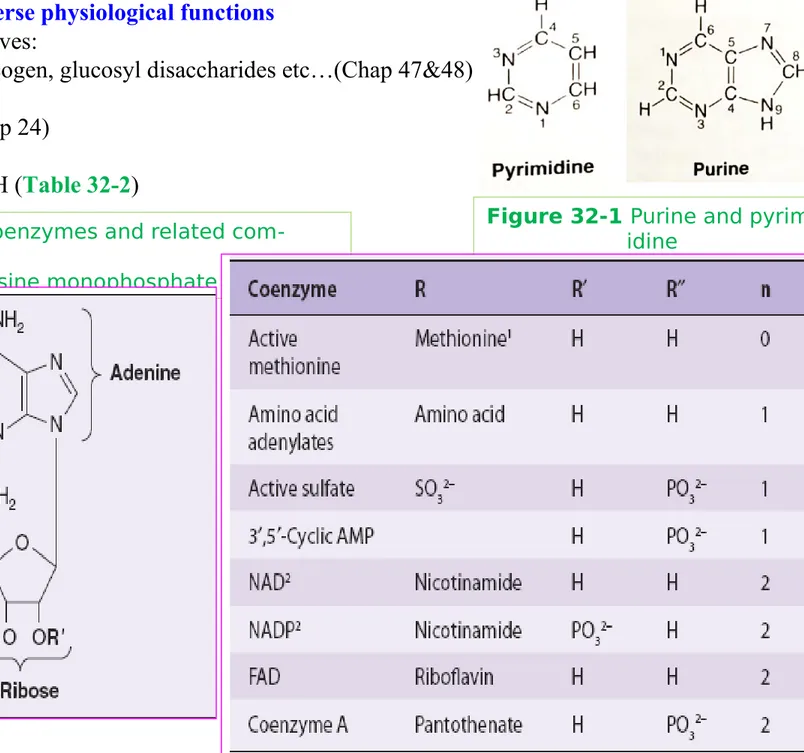
Table. 32-2 Many coenzymes and related
com-pounds are
Derivatives of adenosine monophosphate
Nucleotides serve diverse physiological functions UDP-sugar derivatives:
~ biosynthesis of glycogen, glucosyl disaccharides etc…(Chap 47&48) CTP:
~ lipid synthesis (Chap 24) Coenzymes:
~ NAD, FAD, CoASH (Table 32-2)
Figure 32-1 Purine and
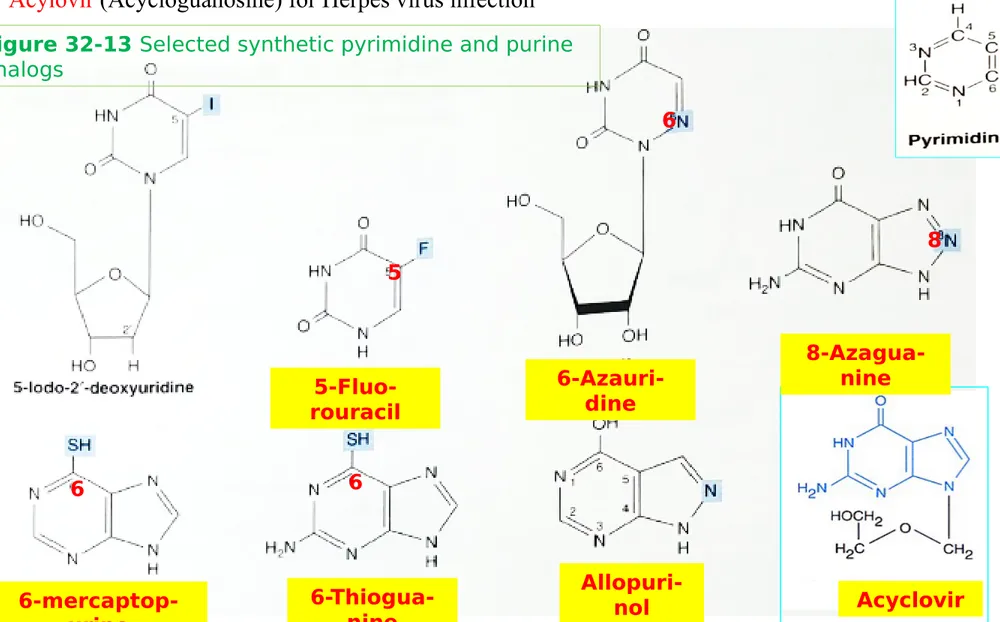
Figure 32-13 Selected synthetic pyrimidine and purine
analogs
Synthetic nucleotide analogs are used in chemotherapy
Synthetic purine & pyrimidine analogs ~ Anti-cancer agents: 5-fluorouracil, 6-thiogua-nine and 6-mercaptopurine, 6-azauridine, 8-azagua6-thiogua-nine incorporated into DNA prior to cell division (Fig. 32-13)
Allopurinol for hyperuricemia and gout
Acylovir (Acycloguanosine) for Herpes virus infection
Allopuri-nol
Figure 32-13 Selected synthetic pyrimidine and purine
analogs 5-Fluo-rouracil Acyclovir 6-Azauri-dine 6-mercaptop-urine 6-Thiogua-nine 8-Azagua-nine 6 6 8 6 5
Figure 32-14 Arabinosylcytosine (cytarabine) and
aza-thioprine
Synthetic nucleotide analogs are used in chemotherapy Synthetic purine & pyrimidine analogs;
~ Cytarabine: chemotherapy of cancer (Fig. 32-14)
Clinical use of drugs
developed by Elion and Hitch-ings
"for their discoveries of important principles for drug treatment"
1988 Nobel Prize in Physiology or Medicine
Purine, Pyrimi-dine base,
nucle-oside Analogs Indications 6-Mercaptopurine, thioguanine Leukemia Azathioprine Organ transplanta-tion, Autoimmune disease Allopurinol Gout
Acyclovir Herpes virus infec-tion
14 Simple precur-sors Base s Nucleo-sides Nucleo-tides DNA, RNA
(Synthetic) Nonhydrolyzable nucleotide triphosphate analogs serve as research tools (Figure 32-15) Synthetic derivatives of nucleoside triphosphates incapable of
undergo-ing hydrolytic release of the terminal phosphoryl group. (Pu/Py, a purine or pyrimidine base; R, ribose or deoxyribose.) Shown are the parent (hydrolysable) nucleoside
triphosphate (top) and the unhydrolyzable β–methylene (center) and γ–imino derivates (bottom).
Synthesis of polynucleotides
DNA & RNA are polynucleotides A nucleoside 5’-triphosphate is added to the 3’-hydroxyl group of a growing peptide chain ~ 5’-phosphoryl group of a mononucleotide (1st) can esterify with 2nd hydroxyl group,
forming a phosphodiester.
~ Phosphodiester bonds link the 3’- and 5’-carbons of adjacent monomers.
~ 2nd hydroxyl group is the 3’-OH of the pentose of the 2nd nucleotide.
forming a dinucleotide (pentose moieties are linked by a 3’,5’-phosphodiester bond to form the “backbone” of RNA and DNA.
The formation of a dinucleotide removes water(H2O) between two mononucleotides
Polynucleotides are directional macromolecules
The base sequence (or primary structure) of a polynucleotide P: phosphodiester bond, a single letter (Bases)
a vertical line; Pentose
5’-end being the one with a free or phosphorylated 5’-hydroxyl.
P A OH P P P T C A pApTpCpA 3’ ATCA 5’ 5’ 3’ 5’ 3’ 5’ 3’ 5’ 3’ 5’ 3’
DNA & RNA are polynucleotides RNAs are far less stable than DNA
2’-hydroxyl group of RNA (absent from DNA: 그림 )
functions as a nucleophile during hydrolysis of the 3’,5’-phosphodiester bond AMP dAMP
그림
2 OH H 2• Under physiologic conditions, the amino and oxo tautomers of purines,
pyrim-idines, and their derivatives predominate.
• Nucleic acids contain, in addition to A, G, C, T, and U, traces of 5-methylcytosine,
5-hydroxymethylcytosine, pseudouridine (tRNA), or N-methylated heterocycles.
• Most nucleosides contain D-ribose or 2-deoxy-D-ribose linked to
N-1
of a
pyrimi-dine or to
N-9
of a purine by a -
glycosidic bond
whose anti- conformers
predomi-nate.
• A primed numeral locates the position of the phosphate on the sugars of
mononu-cleotides (eg, 3'-GMP, 5'-dCMP). Additional phosphoryl groups linked to the first by
acid anhydride bonds
form nucleoside diphosphates and triphosphates.
SUMMARY I
• Nucleoside triphosphates have high group transfer potential and participate in
covalent bond syntheses. The cyclic phosphodiesters cAMP and cGMP function as
intracellular second messengers.
• Mononucleotides linked by 3',5'-phosphodiester bonds form polynucleotides,
direc-tional macromolecules with distinct 3'- and 5'-ends. For pTpGpTp or TGCATCA,
the 5-end is at the left, and all phosphodiester bonds are 3
'-> 5'.
• Synthetic analogs of purine and pyrimidine bases and their derivatives serve as
anticancer drugs
either by inhibiting an enzyme of nucleotide biosynthesis or by
being incorporated into DNA or RNA
SUMMARY II
Nucleotides
2019 년 1 학기Text : Harper’s Biochemistry Refs: Devlin’s, Lippincott’s
Ch. 32. Nucleotides
Biomedical importance:
Biosynthesis is stringently regulated and coordinated by feedback mechanisms
Genetic diseases of purine metabolism: Gout, Lesch-Nyhan syndrome, adenosine deaminase deficiency
•Nucleic acids:
P
hosphate(s) - (d)D-Ribose (sugar) - Base(s)(Purine/Pyrimidine)Purines & pyrimidines are dietarily nonessential: Synthesis from intermediates
Ingested nucleic acids digested to (mononucleotides)bases and Oxidized & excreted in the urine. Injected compounds are incorporated: Injected [3H] thymidine is incorporated into
newly synthesized DNA, thus used to measure the rate of DNA synthesis.
Biosynthesis of purine nucleotides: Growth and Tissue regeneration.
Sources of each atom (Figure 33-1) Isotopic precursors to pigeons.
Three processes for purine nucleotides biosynthesis: Synthesis from amphibolic intermediates; Fig. 33-2, 33-3
Phosphoribosylation; Fig. 33-4
Phosphorylation of purine nucleotides N10
-Formyl-tetrahydrofolate
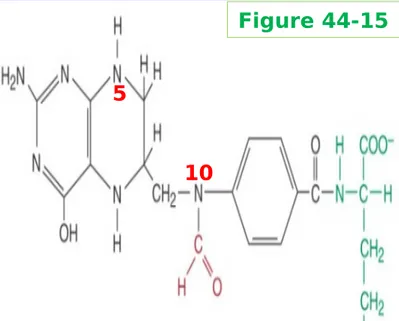
Figure 33-1 Sources of the nitrogen and carbon
atoms
of the purine ring
N5, N10 -Formyl-tetrahydrofolate N10-Formyl-tetrahydro-folate 10 5 Figure 44-15
Amide nitrogen of glutamine Glycine Aspartate
Inosine MonoPhosphate (IMP) is synthesized from amphibolic intermediates
α-D-ribose 5-phosphate (PRPP; PhosphoRibosylPyroPhosphate) IMP (Figure 33-2) PRPP; an intermediate in the purine salvage pathway and in the biosynthesis of pyrimidine
nucleotides, NAD+, and NADP+.
Glutamine, Glycine, N5,N10-Formyl tetrahydrofolate
PRPP
Figure 33-2 Purine biosynthesis from ribose-5-phosphate and
ATP Glutamine Glutamine Glycine N5, N10 -Formyl-tetrahydrofolate
Inosine MonoPhosphate (IMP) is synthesized from amphibolic intermediates α-D-ribose 5-phosphate (PRPP)
Glutamine, glycine, N5,N10-Formyl tetrahydrofolate
Aspartate, N10-Formyl tetrahydrofolate IMP (Figure 33-2)
Aspartate
N10 -Formyl-tetrahydrofolate
IMP
Figure 33-2 Purine biosynthesis from ribose-5-phosphate and
Inosine MonoPhosphate (IMP) AMP & GMP (Figure 33-3) IMP to AMP (ADP ATP)
IMP to XMP GMP (GDP GTP)
Figure 33-3 Conversion of IMP to AMP and GMP
IMP
AMP
GMP
XMP
Multifunctional catalysts participate in purine nucleotide biosynthesis (Figure 33-2) In eukaryotes, multifunctional enzymes
reactions (3,4&6),,,,, (7&8),,,, (10&11)
Antifolate drugs block purine nucleotide biosynthesis (Figure 33-2) Rnxs 4, 10 can be blocked by antifolate drugs
~ Compounds that block purine synthesis by inhibiting formation of folic acid cancer chemotherapy
Figure 33-2 Purine biosynthesis from ribose-5-phosphate and
ATP N5, N10 -Formyl-tetrahydrofolate N10 -Formyl-tetrahydrofolate
IMP
1
0
4
Reaction 2 Reaction 5 Glutamine analogs block purine nucleotide biosynthesis (Figure 33-2)
diazanorleucine (reaction 2) (reaction 13 &14) Azaserine (reaction 5)
Synthetic Purine analogs block purine nucleotide biosynthesis (Figure 33-2/ Fig. 32-13) 6-mercaptopurine (reaction 13 &14)
mycophenolic acid (reaction 14)
IMP
GMP
XMP
AMP
1 4 1 3 Leukemia (Chapa. 32)“Salvage reactions” convert purines & their nucleotides to mononucleotides (Figure 33-2/33-3/ 33-4) “Salvage reaction”:
far less energy than de novo synthesis. 1) Free purines to purine 5’-mononucleotide Pu + PR-PP Pu-RP + Ppi
** Two phosphoribosyl transferases (Figure 33-4) Adenine + PRPP ; AMP (APRTase)
hypoxanthine - guanine + PRPP ; IMP or GMP (HPRTase)
2) Phosphoryl transfer from ATP to nucleoside Pu-R + ATP -> PuR-P + ADP
Liver mainly provides purines & purine nucleosides for salvage and for utilization
A low level of PRPP glutamyl amidotransferase (Fig. 33-2, reaction 2) in human requires exogenous
purines
Brain, erythrocytes, and PMN leukocytes : unable to synthesize 5’-phosphoribosylamine (Fig. 33-2, structure III) and thus, utilize
exogenous purines to form nucleotides.
Figure 33-4 Phosphoribosylation of adenine,
hypoxanthine, and guanine to form AMP, IMP, GMP
PRPP
PRPP
PRPP
Adenine AMP IMP GMP Hypoxanthine GuanineAPRTase
H(G)PRTase
“Salvage reactions” convert purines & their nucleotides to mononucleotides (Figure 33-2/33-3/ 33-4) Lesch-Nyhan Syndrome; an overproduction of hyperuricemia
a defect in hypoxanthine-guanine phosphoryribosyl transferase, an enzyme of a purine salvage (Figure 33-4)
Rise in PRPP results in purine overproduction (Figure 33-2/33-3 )
Figure 33-4 Phosphoribosylation of adenine,
hypoxanthine, and guanine to form AMP, IMP, GMP
PRPP
Figure 33-2 Purine biosynthesis
Hepatic Purine Biosynthesis is stringently regulated (Figure 33-2/33-3/ 33-4) AMP & GMP feedback regulate PRPP glutamyl aminotransferase
Major determinant of purine synthesis: PRPP concentration
Feedback inhibition of rxn 1,2 by AMP, GMP, ADP, GDP (Figure 33-5)
Reactions ① and ② are catalyzed by PRPP synthase and by PRPP glutamyl
amido-transferase, respectively. : Feedback inhibition by intermediates of the pathway
Figure 33-5 Control of the rate of de novo purine nucleotide
biosynthe-sis.
PRPP
AMP IMP GMP ADP GDP ATP GTP Figure 33-2 Figure 33-3 1 1 2 2Hepatic Purine Biosynthesis is stringently regulated (Figure 33-2/33-3/ 33-4) AMP & GMP feedback regulate PRPP glutamyl aminotransferase
AMP & GMP feedback regulate their formation from IMP (Figure 33-6) AMP & GMP feedback inhibition (Rxn 12, 14) (Figure 33-3)
Cross requirement of GTP (Rnx 12) or ATP (Rnx 15) (Figure 33-3) decreases synthesis of one purine nucleotide
when there is a deficiency of the other nucleotide.
Figure 33-6 Regulation of the interconversion of IMP to adenosine
nu-cleotides
and guanosine nucleotides 12 14 Adenylsuccinate synthase
1
2
IMP dehydrogenase1
4
GTP
ATP
1 5 GMP XMP 1 5Figure 33-3
Hepatic Purine Biosynthesis is stringently regulated (Figure 33-2/33-3/ 33-4) AMP & GMP feedback regulate PRPP glutamyl aminotransferase
AMP & GMP feedback regulate their formation from IMP (Figure 33-6) AMP & GMP feedback inhibition (Rxn 12, 14) (Figure 33-3)
AMP & GMP inhibit salvage reaction (Figure 33-4) AMP(GMP) inhibit
Hypoxanthine-guanine phosphoribosyltransferase (H(G)PRTase)
Figure 33-4 Phosphoribosylation of adenine,
hypoxanthine, and guanine to form AMP, IMP, GMP
PRPP
PRPP
PRPP
Adenine AMP IMP GMP Hypoxanthine GuanineReduction of ribonucleotide diphosphates forms deoxyribonucleoside diphosphates (dNDPs ) Reduction of 2'-hydroxyl by ribonucleotide reductase complexes (Fig. 33-7): NDPs → dNDPs Enzyme complex is functional only when cells are actively synthesizing DNA
Reduction requires thioredoxin, thioredoxin reductase, NADPH+H+
Figure 33-7 Reduction of ribonucleoside
Figure 33-8 Regulation of the purine and pyrimidine
ri-bonucleotides to their respective 2’-deoxyriri-bonucleotides
Reduction of ribonucleotide diphosphates forms deoxyri-bonucleoside diphosphates (d-NDPs ) (Figure 33-8) Reduction of ribonucleoside diphosphates (NDPs) to deoxyri-bonucleoside diphosphates (d-NDPs):::
~ Complex regulatory controls based on balanced production of deoxyribonucleotides for DNA synthesis (Figure 33-9 연결 - 참조 ) Purine nucleotides Biosynthesis (Fig. 33-2) Pyrimidine nucleotides Biosynthesis (Fig. 33-9)
Biosynthesis of Pyrimidine Nucleotides
Biosynthetic pathway for pyrimidine nucleotides (Figure 33-9)
1st reaction: cytosolic carbamoyl phosphate synthase II (CPS II) (mitochondrial CPS I of urea cycle;; Figure 28-12)
CO2 + glutamine +ATP -> ->-> UMP -> UDP
CO
2+ glutamine +ATP
1
Figure 33-9 The biosynthetic pathway for pyrimidine nu-cleotidesCarbamoylphosphate
Synthase II(
CPSII
)
Biosynthesis of Pyrimidine Nucleotides
Biosynthetic pathway for pyrimidine nucleotides (Figure 33-9) 1st reaction: cytosolic carbamoyl phosphate synthase II (mitochondrial CPS I of urea cycle;; Figure 28-12)
CO2 + glutamine +ATP -> ->-> UMP -> UDP
CO
2+ glutamine +ATP
1
Figure 33-9 The biosynthetic pathway for pyrimidine
nu-cleotides
Carbamoylphosphate
Synthase II
(CPS II)
Biosynthetic pathway for pyrimidine nucleotides (Figure 33-9)
1st reaction: cytosolic carbamoyl phosphate synthase II (cf, mitochondria CPS I of urea cycle)
CO2 + glutamine +ATP -> ->-> UMP -> UDP
UDP-> UTP-> CTP, UDP-> dUDP-> dUMP-> TMP
Figure 33-9 The biosynthetic pathway for pyrimidine
nu-cleotides
dTMP
Ribonucleotide reductaseUDP
UMP
dUMP
CTP
UTP
(
Fig. 33-7)
d
UDP
Conversion of dUMP to dTMP by thymidylate synthase and
dihydrofolate reductase (Rnx 12) FdUMP
::: inhibition of thymidylate synthase
Figure 32-13
1
2
dUMP
dTMP
Thymidylate synthase D ih yd ro fo la te re du ct as e Methotrexate Figure 33-9-FdUMP
-5-fluorouracil H Uracil
Folic acid
Methotrexate
Conversion of dUMP to dTMP by thymidylate synthase and dihydrofo-late reductase (Rnx 12)
Methotrexate (anticancer drug;) ::: inhibitor of dihydrofolate reductase
1
2
dUMP
dTMP
Thymidylate synthase D ih yd ro fo la te re d u ct as e Methotrexate Figure 33-9-FdUMP
- Allopuri-nol Certain pyrimidine analogs are substrates for enzymes of pyrimidine nucleotide biosynthesis Orotate phosphoribosyltransferase (rxn 5, Fig. 33-9) converts the drug allopurinol (Fig, 32-13) to nucleotide in which ribosyl phosphate is attached to N-1 of the pyrimidine ring.
Orotate phosphoribosyltransferase (rxn 5, Fig. 33-9) phosphoribosylates the anticancer drug
5-fluorouracil nucleotide
5
Figure 33-9 The biosynthetic pathway for pyrimidine
nu-cleotides
Orotate Phosphoribosyl
transferase rouracil
5-fluo-5 Carbamoylphosphate Synthase II (CPS II) Fig, 32-13 Fig, 32-13
The deoxyribonucleosides of Uracil & Cytosine are salvaged
Salvage reactions converts
~ pyrimidine ribonucleosides uridine & cytidine nucleotides (UMP, UDP, UTP….CMP, CDP, CTP) ~ pyrimidine deoxyribonucleosides thymidine & deoxycytidine nucleotides (dTMP….dCMP…) ATP-dependent phosphoribosyltransferases (kinases) catalyze
~ 2’-deoxy-(cytidine/ guanosine/ adenosine) their corresponding nucleoside triphosphates (dCTP,, dGTP,, dATP)
Orotate phosphoribosyltransferase (rxn 5, Fig. 33-9), an enzyme of pyrimidine nucleotide synthesis, salvages orotic acid by converting it to orotidine monophosphate (OMP)
5
Figure 33-9 The biosynthetic pathway for pyrimidine
nu-cleotides
Orotate Phosphoribosyl
transferase Orotic acid
(OA) OMP
Regulation of Pyrimidine Nucleotide Biosynthesis
Gene expression & enzyme activity both are regulated
(
Fig. 33-9): allosteric regulation Rxn 1 (CPSII) is inhibited by UTP but activated by PRPP
Rxn 2 (ATCase) is inhibited by CTP but activated by ATP (Fig. 33-10)
CO2 + glutamine +ATP
Figure 33-9 The biosynthetic pathway for pyrimidine
nu-cleotides Carbamoylphosphate Synthase II 1 2 (-)UTP (+)PRPP Aspartate transcarbamoylase (-)CTP (+)ATP
Purine & pyrimidine nucleotides biosynthesis are coordinately regulated
Cross regulation: PRPP synthase (rxn , Fig. 33-2) is inhibited by both pu and py nucleotides
Figure 33-8 Regulation of the purine and pyrimidine
ri-bonucleotides to their respective 2’-deoxyriri-bonucleotides
P ur in e nu cl eo ti de s B io sy nt he si s ( F ig . 3 3-2 ) P yr im id in e nu cl eo ti de s B io sy nt he si s ( F ig . 3 3-9 ) 1 1
Regulatory aspects of purine & pyrimidine nu-cleotides reduction to their respective 2’-de-oxyribonucleotide (Fig. 33-10)
Humans Catabolize Purines to Uric acid ~ Adenosine and Guanosine
uric acid in human (Fig. 33-11)
Uricase converts uric acid to allantoin (wa-ter-soluble). ~~ Since humans lack uricase, end product of purine catabolism is uric acid.
Uric acid Guanosine Adenosine deaminase Inosine Adenosine Guanine Hypoxanthine Xanthine
Figure 33-11 Formation of uric acid from purine nucleo -sides
Gout is a Metabolic Disorder of Purine Catabolism Defect in PRPP synthase : Vmax (rxn 1, Fig 33-2) ~ Overproduction and overexcretion of purine catabolites When serum urate levels exceed the solubility limit, ~ Sodium urate crystallizes,
~ causing inflammatory reaction, gouty arthritis (통풍관절 염 )
Most cases of gout: abnomalities in renal handling of uric acid. ③ Treatment: Allopurinol (Fig. 33-11 참조 )
PRPP
PRPP synthase Xanthine oxidase XanthineUric acid
Xanthine oxidaseOther Disorders of Purine Catabolism
: Genetic disorders of purine catabolisms
Lesch-Nyhan syndrome
~ Hyperuricemia
~ Defect in HGPRTase (Fig. 33-4)
rise in PRPP purine overproduction (Fig. 33-2)
Von Gierke's disease
~ Glucose-6-phosphatase deficiency
increase in ribose 5-phosphate (Fig. 33-2) purine overproduction & hyperuricemia
Hypouricemia
(increased excretion of hypoxanthine and xanthine; Fig. 33-11) ~ Xanthine oxidase deficiency
due to a genetic defect or to severe liver damages Xanthinuria
PRPP
IMP Hypoxanthine Hypoxanthine-guanine Phosphoribosyltransferase (H(G)PRTase)PRPP
Uric acid Guanine Hypoxanthine Xanthine Xanthine oxidaseFig. 33-2
Adenosine deaminase Inosine Adenosine Guanosine Uric acid Guanine Hypoxanthine Xanthine
Other Disorders of Purine Catabolism
Adenosine deaminase deficiency (Fig. 33-11) ~ an immunodeficiency disease
T&B cells are sparse and dysfunctional
Purine nucleoside phosphorylase deficiency(Fig. 33-11) (InosineHypoxnathine; GuanosineGuanine)
~ severe deficiency of T cells with normal B cell function Immune dysfucntions:
result from accumulation of dATP & dGTP, Inhibit ribonucleotide reductase, (Fig. 33-9) Thereby deplete cells of DNA precursors
Purine nucleoside phosphorylase Ribonucleotide reductase UDP dUDP dUMP dTMP
Catabolism of Pyrimidines Produces
Water-soluble Metabolites
β-ureidopropionase (from pyrimidine precursors)
(Fig. 33-12)
~ water-soluble end product of pyrimidine catabolism
CO
2, NH3, β-alanine, and β-aminoisobutyrate~ β-aminoisobutyrate forms succinyl-CoA
(Fig. 20-2)
~ Excretion of β-aminoisobutyrate increases
in leukemia & severe x-ray radiation exposure due to increased destruction of DNA
~ β-hydroxybutyric aciduria;
deficiency in dihydropyrimidine dehydrogenase ~ uraciluria-thyminuria
pyrimidine catabolism disorder
Cytosine Dihydropyrimidine dehydrogenase -Ureidopropionase -Alanine -Amidoisobutyrate CO2 + NH3
Fig. 33-12
Uracil ThymineOverproduction of pyrimidine catabolites is only
rarely associated with clinically significant
ab-normalities
( vs abnormal purine catabolisms)
Since end-products of pyrimidine catabolism are highly water-soluble, pyrimidine overproduction results in few clinical signs or symptoms
Orotic acidurias ~ Reye syndrom;
severely damaged mitochondria to utilize carbamoyl phosphate
(Fig. 28-12; def.mt CPS1; urea cycle),
cytosolic overproduction of orotic acid (Fig. 33-9).
*** Type I (Rnx 5/6), II (Rnx 6)-orotic aciduria ~ Deficiency of a urea cycle enzyme:
mt ornithine transcarbamoylase deficiency (Fig. 28-12) CAP(cytosol) increase
results in excretion of pyrimidine precursors orotic aciduria